
McGill Univ. Libraries and the generosity of Dr. Richard L. Tomlinson
 |
Brought to you by: McGill Univ. Libraries and the generosity of Dr. Richard L. Tomlinson |
|||
|
|
|
| Leukemia
Research Volume 29, Issue 5, May 2005, Pages 565-572 |
| ||||
|
|
| ||||
Cytokinetics and mechanism of action of AKO4: a novel nitrogen mustard targeted to bcr-abl
Athanasia Katsoulasa,
Zakaria Rachida,
Fouad Brahimia,
James McNameeb
and Bertrand J. Jean-Claudea,
![]() ,
, ![]()
Abstract
The “combi-targeting” concept seeks to design molecules to not only block tyrosine kinase (TK) activity but also to induce DNA damage. Here we design AK04, a molecule that combines the pharmacophore chlorambucil with that of STI-571 (Gleevec™). The results showed that although a less potent abl TK inhibitor than STI571, AK04 was capable of significantly blocking bcr-abl phosphorylation not only in a purified abl assay but also in the bcr-abl+ K562 cells. In contrast to STI571 and like chlorambucil, it induced a dose-dependent increase in DNA damage in these cells. More importantly, AK04 was 12–32-fold more potent than chlorambucil in all bcr-abl+ cells of our cell panel. In the isogenic human megakaryocytic Mo7e and Mo7/bcr-abl cells, AK04 selectively killed the bcr-abl transfectants. Flow cytometry revealed that despite being a five-fold less potent inhibitor of bcr-abl than STI-571, it induced a significant dose-dependent increase in levels of cell death by apoptosis in KU812 cells 24 h post-treatment. Under these conditions, chlorambucil did not induce any significant level of apoptosis. These results suggest that AK04 is a nitrogen mustard with binary bcr-abl/DNA targeting effects, a property that may account for its superior potency when compared with the classical mustard chlorambucil.
Keywords: CML; Chlorambucil; Imatinib; Apoptosis;
Double-strand breaks
Article Outline
1. Introduction
The nitrogen mustards have been used in the chemotherapy of cancer for over
50 years. Their mechanism of action is based on their ability to induce covalent
binding to DNA and especially by bifunctional reaction to create DNA
cross-linking adducts in tumour cells [1].
The principal site of alkylation on DNA is the N7-position of guanine and this
is often followed by reaction with N3-adenine or another N7-guanine at the
opposite DNA strand [2],
[3],
[4]
and [5].
These lesions are believed to be responsible for the significant cytotoxicity of
nitrogen mustards. More recently, their conjugation to DNA-reading peptides was
shown to target their alkylation reaction to adenine-rich DNA sequences and to
confer them higher levels of cytotoxicity in tumour cells when compared with
their non-conjugated counterparts [6],
[7]
and [8].
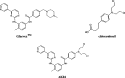 |
Display Full Size version of this image (7K) |
The stability of nitrogen mustards under physiological conditions have been improved by substitution of the methyl group with a phenyl ring to reduce the electron density at the nitrogen, leading to stable molecules termed aniline mustards. One such agent chlorambucil is one of the most potent alkylators used in the therapy of chronic lymphocytic leukemia (CLL) [9]. Despite the extensive use of aniline mustards in the therapy of various leukemias, little effort has been directed towards targeting them to leukemia-specific tumour markers. The bcr-abl oncogene being the proven cause of chronic myelogenous leukemia (CML) [10], [11] and [12], we considered it to be an attractive marker for the first attempt to target aniline mustard to a form of leukemia. More importantly the bcr-abl oncogene is associated with chemoresistance to cytotoxic agents through DNA repair and its anti-apoptotic function [13] and [14].
One of the most potent inhibitors of the abl tyrosine kinase is STI571 (Gleevec), a 2-phenylaminopyrimidine that competitively binds to the ATP site of its kinase domain, thereby blocking its phosphorylation activity [15], [16] and [17]. This molecule has proven one of the most effective agents in the clinic in the therapy of CML [16]. However, due to acquired resistance associated with the repeated dosing schedule of the drug, and the short duration of response in patients in the blast crisis, interest is now turned towards combining the latter drug with classical cytotoxic agents [11] and [18]. Here we designed a novel strategy that seeks to combine both bcr-abl inhibitory and cytotoxic functions within one single molecule, reducing the pharmacotoxicology associated with these two antiproliferative mechanisms to that of a single molecule. The feasibility and efficacy of this approach termed “combi-targeting” has now been demonstrated [19], [20], [21] and [22].
The design was based on replacing the methylpiperazine ring of STI571 by a nitrogen mustard function fused with a benzamide spacer, leading to the “2-phenylaminopyrimidine aniline mustard conjugate termed “AKO4”. Here we report on the cytotoxicity and the mechanism of action of the compound in vitro in comparison with that of the aniline mustard chlorambucil.
2.1. Drug treatment
Chlorambucil was purchased from Sigma-Aldrich. AK04 was synthesized in our laboratory by direct coupling of 2-(3-aminophenylaminopyrimidinopyridine with 4-(bis-chloroethyl)aminobenzoyl chloride. In all assays, drug was dissolved in DMSO and subsequently diluted in RPMI-1640 containing 10% fetal bovine serum (Wisent Inc., St-Bruno, Canada) immediately before the treatment of cell cultures. In all assays, the concentration of DMSO never exceeded 0.2% (v/v).
2.2. Cell culture
K562 (American Type Culture Collection, Manassas, VA), KU812 and U937 cells (generous gifts from Dr. Carlo Gambacorti-Passerini, Division of Experimental Oncology D and Medical Oncology C, Tumour National Institute, Italy) were maintained in RPMI-1640 supplemented with 10% FBS and antibiotics as described previously [19]. Mo7e and Mo7p210 [27] (generous gift from Dr. Brian J. Druker, Division of Hematology and Medical Oncology, Oregon Health Sciences, USA) were grown in RPMI-1640 supplemented with 10% FBS, 2 % (v/v) l-glutamine, 10 ng/ml granulocyte-macrophage colony stimulating factor (GM-CSF) and antibiotic. All cells were maintained in an atmosphere of 5% CO2.
2.3. Kinase assay
This assay is similar to the EGFR kinase one with minor modifications [19] and [23]. Briefly, Nunc Maxisorp 96-well plates were incubated overnight at 37 °C with 100 μl/well of 0.25 mg/ml PGT in PBS. Excess PGT was removed and the plate was washed three times with wash buffer Tween 20 (0.1%) in PBS. The kinase reaction was performed by using 1.0 ng/well c-abl (Bioscience Inc., California). The compound was added and phosphorylation initiated by the addition of ATP (50 μM). After 10 min at room temperature with constant shaking, the reaction was terminated by the addition of 50 μl of 250 mM EDTA prior to aspiration and by rinsing the plate four times with wash buffer. Phosphorylated PGT was detected following a 25 min incubation with 50 μl/well of HRP-conjugated PY20 anti-phosphotyrosine antibody (Santa Cruz Biotechnology, California) diluted to 0.2 μg/ml in blocking buffer (3% bovine serum albumin; 0.05% Tween 20 in PBS). Antibody was removed by aspiration, and the plate washed four times with wash buffer. The signals were developed by the addition of 50 μl/well of 3,3′,5,5′-tetramethylbenzidine peroxidase substrate (Kirkegaard and Perry Laboratories, Gaithersberg, MD, USA) and following blue color development, 50 μl of H2SO4 (0.09 M) was added per well, and plates were read at 450 nm using a Bio-Rad ELISA reader (model 2550).
2.4. The bcr-abl autophosphorylation assay
Inhibition of receptor autophosphorylation in viable cells was determined by anti-phosphotyrosine Western blots. Cells were plated at a concentration of 2 × 106 cells per well and incubated with the compounds for 2 h. Thereafter, they were washed with PBS and re-suspended in cold lysis buffer [50 mM Tris–HCl pH 7.5; 150 mM NaCl; 1% Nonidet P-40, 1 mM EDTA; 5 mM NaF; 1 mM Na3VO4; protease inhibitor tablet (Roche Biochemicals, Laval, Canada)]. The lysates were kept on ice for 30 min and collected by centrifugation at 10,000 rpm for 20 min at 4 °C. Protein concentrations were determined against a standardized control using the Bio-Rad protein assay kit (Bio-Rad Laboratories, Hercules, CA). Equal amounts of protein from each cell lysate were added to an 8% SDS-polyacrylamide gel electrophoresis and transferred to a polyvinylidene difluoride membrane (Millipore, Bedford, MA). Non-specific binding on the membrane was minimized with a blocking buffer containing nonfat dry milk (3%) in PBS. The membrane was blotted for 1 h with anti-phosphotyrosine antibody 4G10 (Upstate Biotechnology, New York) or anti-abl antibodies (Upstate Biotechnology, New York). It was subsequently incubated with HRP-goat anti-mouse antibody (Bio-Rad laboratories) or with HRP-goat anti-rabbit (Upstate Biotechnology, New York) and the bands visualized with an enhanced chemiluminescence system (Amersham pharmacia Biotech, Buckinghamshire, UK). Band intensities were measured using the SynGene GeneTools software package.
2.5. MTT assay
The cytotoxic effects of our compounds were evaluated using the 3-(4,5-dimethylthiazo-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay [24] with minor modification. Briefly, cells were grown in 12-well plates and then exposed to compounds for 96 h. MTT (50 μl) of (5 mg/ml in sterile PBS) was added to 1 ml of media and the plates were incubated for 2–3 h at 37 °C. The resulting colored residue was dissolved in DMSO and optical density was read for each well at 570 nm using a Bio-Rad microplate reader (model 2550). Each point represents the average of at least two independent experiments run in duplicate.
2.6. Alkaline comet assay for quantitation of DNA damage
The alkaline comet assay was performed as previously described [6]. The cells were exposed to drugs for 2 h, and resuspended in PBS. Cell suspensions were diluted to approximately 106 cells, and mixed with agarose (1%) in PBS at 37 °C in a 1:10 dilution. The gels were cast on Gelbond strips (Mandel Scientific, Guelph, Canada) using gel casting chambers and then immediately placed into a lysis buffer [2.5 M NaCl, 0.1 M tetra-sodium EDTA, 10 mM Tris–base and 1% (v/v) Triton X-100, pH 10.0]. After being kept on ice for 30 min, the gels were gently rinsed with distilled water and immersed in a second lysis buffer (2.5 M NaCl, 0.1 M tetra-sodium EDTA, 10 mM Tris–base) containing 1 mg/ml proteinase K for 60 min at 37 °C. Thereafter, the gels were rinsed with distilled water, incubated in alkaline electrophoresis buffer for 30 min at 37 °C, and electrophoresed at 19 V for 20 min. The gels were subsequently rinsed with distilled water and placed in 1 M ammonium acetate for 30 min. Thereafter, they were soaked in 100% ethanol for 2 h, dried overnight, and stained with SYBR Gold (1/10,000 dilution of stock supplied from Molecular Probes, Eugene, OR) for 20 min. Comets were visualized at 330× magnification and DNA damage was quantitated using the tail moment parameter (i.e., the distance between the barycenter of the head and the tail of the comet multiplied by the percentage of DNA within the tail of the comet). A minimum of 50 cell comets were analyzed for each sample, using ALKOMET version 3.1 image analysis software.
3.1. Design of AK04
X-ray structure of c-abl co-crystallized with an analogue of STI571 demonstrated that in addition to hydrogen bonds between Met 318 amide backbone and the pyridine nitrogen, between the pyrimidine NH and the Thr 315 OH group, the amide bond of the benzamido moiety was involved in a donor–acceptor interaction wherein the oxygen of Glu 286 formed a hydrogen bond with the benzamide NH. The oxygen of the latter accepts a hydrogen bond from the amide backbone of Asp 381. The pyrazinyl moiety is located in a partially hydrophobic pocket forming Van der Waals interactions with Val 289, Phe 359 and Asp-381. Since previous studies demonstrated that some binding affinity could be retained if the phenylaminopyrimidine moiety remained intact [25], we decided to modify the benzamido extension in a manner that preserves the amido NH. Further, the nitrogen mustard function was appended to the 4-position in order to straddle the 4-methylpiperazino group of STI-571. Since our aniline mustard group does not occlude the partially hydrophobic pocket observed in the abl X-ray structure [17] as well as the piperazine ring, we did not expect the resulting conjugate AK04 to possess tyrosine kinase inhibitory activities superior to that of STI-571 (Scheme 1). Affinity in the submicromolar level would be sufficient to target our aniline mustard to bcr-abl. Thus, we expected this compound to have a binary targeting property: (a) ability to block tyrosine kinase via its N-benzoylaminophenylpyrimidine system, (b) ability to damage DNA via its mustard appendage.
Display Full Size version of this image (20K) Scheme 1. Overlay model of AK04 and ST1571. One of the chloroethyl groups increases the steric bulk of 4-position of the benzamide moiety.
3.2. The bcr-abl inhibition in a whole cell assay
3.2.1. Enzyme assay
Binding affinity using an ELISA that utilizes polyglutamate tyrosine (4:1) as a substrate [19] showed an IC50 of 0.22 μM, an affinity that is approximately 5-fold less than that of STI571 (IC50 = 0.04 μM) (Fig. 1). As expected, chlorambucil a butyric acid tethered to an aniline mustard did not show any significant v-abl tyrosine kinase inhibitory activity (IC50 > 100 μM).
Display Full Size version of this image (5K) Fig. 1. Inhibition of purified ABL TK activity by AK04 in comparison with STI571 in a competitive ABL binding assay. Phosphorylation of a PGT substrate by purified ABL was measured using immunoassay with anti-phosphotyrosine antibodies.
3.2.2. Whole cell assay
While inhibition of bcr-abl in an ELISA is indicative of the affinity of the inhibitor for its cognate site, because of the high intracellular ATP concentrations, the inhibition of bcr-abl autophosphorylation in a whole cell assay remains the ultimate test. Thus, we compared the ability of the three drugs to block bcr-abl activation in K562 cells using a Western blotting assay as described previously [26]. The results showed that like STI571, AK04 blocked bcr-abl phosphorylation in a dose-dependent manner without affecting the levels of bcr-abl (Fig. 2a). In contrast, no bcr-abl activity was seen for the DNA alkylating aniline mustard chlorambucil (Fig. 2c).
Display Full Size version of this image (50K) Fig. 2. Inhibition of tyrosine phosphorylation of bcr-abl in K562 cells. Cells were incubated for 2 h in the presence of the indicated concentration of AK04 (a), STI571 (b) and chlorambucil (c). They were lysed and equal amounts of lysate were analyzed by immunoblotting with antiphosphotyrosine or anti-abl antibodies.
3.3. Flow cytometric analysis of apoptosis
KU812 cells exposed to STI571 and AK04 showed a dose-dependent increase in a sub-G1 peak, indicating induction of apoptosis. Levels of cell death by apoptosis induced by AKO4 were approximately 1.3–1.6-fold less than those by STI571 (Fig. 3). In contrast, chlorambucil did not induce any discrete sub-G1 peak over the whole dose range, indicating that its mechanism of action is markedly different from that of AKO4 or STI571 (Fig. 3).
Display Full Size version of this image (54K) Fig. 3. Cell cycle analysis following drug treatment in KU812 cells (bcr-abl+). Cells were treated with of STI571, AK04 and chlorambucil, or were untreated controls. After 24 h, cell cycle analysis was performed using propidium iodide staining and flow cytometry as described in Section 2.
3.4. DNA damage
AK04 being a nitrogen mustard, its ability to damage DNA was tested in comparison with chlorambucil. DNA strand breaks were detected by the comet assay following a 2 h drug exposure. As shown in Fig. 4, AK04 induced a dose-dependent increase in DNA damage in K562 cells although its DNA damaging potential appeared approximately two-fold weaker than that of chlorambucil. Thus, AK04 can be considered a nitrogen mustard with a rather mild alkylating property.
Display Full Size version of this image (4K) Fig. 4. Quantitation of DNA damage using the alkaline comet assay. DNA damage induced by AKO4 and chlorambucil in the K562 cell line. Tail moment was used as a parameter for the detection of DNA damage in K562 cells exposed to AKO4 and chlorambucil for 2 h.
3.5. Cytotoxicity assay
To test whether the binary bcr-abl/DNA targeting properties of AK04 translate into selective potency in bcr-abl+ cells, an MTT assay was used with the purpose of comparing the effects of the different drugs on K562, KU812, U937 and an isogenic megakaryocytic pair of cells (Mo7e and Mo7/p210) (Fig. 5). The cells were exposed to drugs continuously for 4 days. In contrast to chlorambucil, AK04 showed selective potency for bcr-abl expressing cells with IC50 values in the 2.5–6 μM range (Table 1). More importantly, in the isogenic pair of cell line, AK04 induced five-fold higher level of cell-killing in the bcr-abl transfectant when compared with its wild type counterpart (Fig. 6a). In contrast, chlorambucil showed significant selectivity for the normal cells, Mo7e (Fig. 6b). This is again indicative of a marked difference between the mechanisms of action of these two nitrogen mustards.
Display Full Size version of this image (8K) Fig. 5. Cytotoxic effects of AK04 (a) and chlorambucil (b) in K562 and U937 cells. Cells were exposed to each drug for 96 h. Cell growth was measured using the MTT assay. Each point represents two independent experiments in triplicate.
Table 1.Comparison between the cytotoxicity of the combi-molecule AKO4 and that of the classical nitrogen mustard chlorambucil against leukemia cells with varied bcr-abl expression status
a IC50 values are the mean of two independent experiments.
Cell lines bcr-abl IC50 (μM)a
AKO4
Chlorambucil
K562 + 2.46 79.04 KU812 + 2.89 36.99 U937 − 160.6 36.16 Mo7/p210 + 5.70 10.99 Mo7e − 25.35 1.26
Display Full Size version of this image (10K) Fig. 6. Cytotoxic effects of (a) AK04 and (b) chlorambucil in Mo7e and Mo7/p210 cells. Cells were exposed to each drug for 96 h. Cell growth was measured using the MTT assay. Each point represents two independent experiments in triplicate.
4. Discussion
The crystal structure of the kinase domain of c-abl in complex with STI571 is now available [15] and [17]. The compound binds to the ATP site of a specific inactive conformation of c-abl in which the loop mimics bound peptide substrates. The 2-phenylaminopyrimidine moiety occludes the region where adenine normally binds, accepting a hydrogen bond at its pyridine nitrogen from Met 318 amide hydrogen. The piperazine binds to a partially hydrophobic pocket forming Van der Waals interactions with valine 289, phe359 and asp 381. The reduced activity of our conjugate AK04 when compared with STI571 may be due to the high degree of freedom of the bulky mustard group when compared with the piperazine ring. Indeed, as evidenced by the overlay model (Scheme 1) one of the chloroethyl groups points away from the 6-membered ring of STI571. Nevertheless, despite the absence of optimal hydrophobic interactions, the appendage of the nitrogen mustard group was not significantly deleterious to binding affinity. The conjugate showed ability to block substrate phosphorylation by bcr-abl both in an enzyme and a whole cell assay. Indeed, AK04 induced 50% inhibition of bcr-abl autophosphorylation at concentrations lower than 12 μM. As expected chlorambucil was inactive in these assays. Thus, AK04 is the first ever reported aniline mustard capable of depleting bcr-abl inhibitory activity in a dose-dependent manner.
AK04 was designed to behave as a combination of a bcr-abl kinase inhibitor like STI571 and a DNA damaging agent like chlorambucil. As demonstrated by the comet assay, it is capable of inducing a dose-dependent increase in strand breaks although with a lesser potency than chlorambucil. This may well be due to the difference of reactivity between the nitrogen mustard function of AK04, when compared with that of chlorambucil. The para-carbonyl of the aniline moiety of AK04 may decrease the electron density at the nitrogen to a level that reduces its alkylating power. Indeed, strong DNA alkylating activity with benzamide mustards are only seen when they are tethered to DNA reading peptide that enhances their delivery to specific DNA sequences [6], [28] and [29].
Despite being a moderate alkylator and a five-fold weaker bcr-abl inhibitor than STI-571, AK04 could induce significant levels of apoptosis in KU812 cells. Perhaps, in contrast to STI-571, its bcr-abl TK inhibitory activity is not the sole property responsible of its ability to induce apoptosis. Its additional ability to damage DNA (although weakly) may at least partially contribute to the enhancement of apoptotic response. Blockade of bcr-abl activity may deplete the levels of DNA repair activity required for the repair of the mustard-induced lesions. A similar mechanism was recently reported to account for the synergistic interaction between chlorambucil and STI-571 in CLL cells [30].
Another critical observation was the inverse selectivity of chlorambucil for the normal cells when compared with AK04 in the isogenic pair of cell lines. It is now known that bcr-abl expression is associated with elevated DNA repair activity through expression of Rad51 [13] and [14]. Thus, in contrast to the wild type cells, DNA repair and antiapoptotic effects in Mo7/p210 may be responsible for resistance to chlorambucil, thereby exacerbating the differential response between the two cell types. Conversely, the selectivity of AK04 can be explained by its ability to block bcr-abl while inducing significant DNA damage, a mechanism that enhances cell killing in the bcr-abl transfectant.
In conclusion, this study demonstrated that by conferring a binary
bcr-abl/DNA targeting property to an analogue of STI571, significantly selective
cell killing could be achieved. Further studies are ongoing in our laboratory to
enhance the bcr-abl inhibitory activity and the DNA damaging potential of AK04
prior to demonstrate its efficacy in vivo.
References
[1] C.J. O’Connor, W.A. Denny, J. Fan, L. Gravatt, B.A. Grigor and D.J. McLennan, Hydrolysis and alkylating reactivity of aromatic nitrogen mustards, J Chem Soc 12 (1991), pp. 1933–1939. Full Text via CrossRef
[2] M.A. Alaoui-Jamali, L. Panasci, G.M. Centurioni, R. Schecter, S. Lehnert and G. Batist, Nitrogen mustard–DNA interaction in melphalan-resistant mammary carcinoma cells with elevated intracellular glutathione and glutathione-S-transferase activity, Cancer Chem Pharmacol 30 (1992), pp. 341–347. Full Text via CrossRef | View Record in Scopus | Cited By in Scopus (16)
[3] R. Geleziunas, A. McQuillan, A. Malapetsa, M. Hutchinson, D. Kopriva and M.A. Wainberg et al., Increased DNA synthesis and repair-enzyme expression in lymphocytes from patients with chronic lymphocytic leukemia resistant to nitrogen mustards, J Natl Cancer Inst 83 (1991), pp. 557–564. View Record in Scopus | Cited By in Scopus (44)
[4] B.B. Bank, Studies of chlorambucil-DNA adducts, Biochem Pharmacol 44 (1992), pp. 571–575. Abstract | Abstract + References | PDF (622 K) | View Record in Scopus | Cited By in Scopus (7)
[5] B.B. Bank, D. Kanganis, F.L. Leonard and R. Silber, Chlorambucil pharmacokinetics and DNA binding in chronic lymphocytic leukemia lymphocytes, Cancer Res 49 (1989), pp. 554–559. View Record in Scopus | Cited By in Scopus (18)
[6] P.G. Baraldi, P. Cozzi, C. Geroni, N. Mongelli, R. Romagnoli and G. Spalluto, Novel benzoyl nitrogen mustard derivatives of pyrazole analogues of distamycin A: synthesis and antileukemic activity, Bioorg Med Chem 7 (1999), pp. 251–262. SummaryPlus | Full Text + Links | PDF (351 K) | View Record in Scopus | Cited By in Scopus (18)
[7] P.G. Baraldi, I. Beria, B. Cacciari, L. Capolongo, P. Cozzi and N. Mongelli et al., Structure–activity relationship of novel tallismustine derivatives: synthesis and antitumor activity, Bioorg Med Chem Lett 6 (1996), pp. 1247–1252. SummaryPlus | Full Text + Links | PDF (205 K) | View Record in Scopus | Cited By in Scopus (22)
[8] P.G. Baraldi, I. Beria, B. Cacciari, P. Cozzi, C. Franzetti and N. Mongelli et al., Synthesis and antitumor activity of novel distamycin derivatives, Bioorg Med Chem Lett 6 (1996), pp. 1241–1246. SummaryPlus | Full Text + Links | PDF (220 K) | View Record in Scopus | Cited By in Scopus (17)
[9] E.Z. Ezdinli and L. Stutsman, Chlorambucil therapy for lymphomas and chronic lymphocytic leukemia, J Am Med Assoc 191 (1965), pp. 444–450.
[10] M.W. Deininger, Vieira SAD, Y. Parada, L. Banerji, EW-F. Lam and G. Peters et al., Direct relation between Bcr-Abl tyrosine kinase activity and Cyclin D2 expression in lymphoblasts, Cancer Res 61 (2001), pp. 8005–8013. View Record in Scopus | Cited By in Scopus (27)
[11] Y. Kano, M. Akutsu, S. Tsunoda, H. Mano, Y. Sato and Y. Honma et al., In vitro cytotoxic effects of a tyrosine kinase inhibitor STI571 in combination with commonly used antileukemic agents, Blood 97 (2001), pp. 1999–2007. Full Text via CrossRef | View Record in Scopus | Cited By in Scopus (165)
[12] D. Wisniewski, C.L. Lambeck, C. Liu, A. Strife, D.R. Veach and B. Nagar et al., Characterization of potent inhibitors of the Bcr-Abl and the c-kit receptor tyrosine kinases, Cancer Res 62 (2002), pp. 4244–4255. View Record in Scopus | Cited By in Scopus (91)
[13] P.Q. Dou, T.F. Mcguire, Y. Peng and B. An, Proteasome inhibition leads to significant reduction of Bcr-abl expression and subsequent induction of apoptosis in K562 human chronic myelogenous leukemia cells, J Pharm Exp Ther 289 (1999), pp. 781–790.
[14] J.F. Dorsey, R. Jove, A.J. Kraker and J. Wu, The pyrido[2,3-d]pyrimidine derivative PD180970 inhibits p210Bcr-Abl tyrosine kinase and induces apoptosis of K562 leukemic cells, Cancer Res 60 (2000), pp. 3127–3131. View Record in Scopus | Cited By in Scopus (85)
[15] T. Schindler, W. Bornmann, P. Pellicena, W.T. Miller, B. Clarkson and J. Juriyan, Structural mechanism for STI-571 inhibition of abelson tyrosine kinase, Science 289 (2000), pp. 1938–1942. Full Text via CrossRef | View Record in Scopus | Cited By in Scopus (622)
[16] M.J. Mauro, M. O’Dwyer, M.C. Heinrich and B.J. Druker, STI571: a paradigm of new agents for cancer therapeutics, J Clin Oncol 20 (2002), pp. 325–334. Full Text via CrossRef | View Record in Scopus | Cited By in Scopus (128)
[17] B. Nagar, W.G. Bornmann, P. Pellicena, T. Schindler, D.R. Veach and T.D. Miller et al., Crystal structures of the kinase domain of c-abl in complex with the small molecule inhibitors PD173955 and Imatinib (STI-571), Cancer Res 62 (2002), pp. 4236–4243. View Record in Scopus | Cited By in Scopus (272)
[18] J.T. Thiesing, J. Ohno-Jones, K.S. Kolibaba and B.J. Druker, Efficacy of STI571, an Abl tyrosine kinase inhibitor, in conjunction with other antileukemic agents against Bcr-Abl-positive cells, Blood 96 (2000), pp. 3195–3199. View Record in Scopus | Cited By in Scopus (252)
[19] F. Brahimi, S. Matheson, J. McNamee, A. Tari and B.J. Jean-Claude, Inhibition of epidermal growth factor receptor-mediated signaling by “combi-triazene” BJ2000, a new probe for the combi-targeting postulates, J Pharm Exp Ther 303 (2002), pp. 238–246. Full Text via CrossRef | View Record in Scopus | Cited By in Scopus (22)
[20] S.L. Matheson, J. McNamee and B.J. Jean-Claude, Differential responses of EGFR−/−AGT-expressing cells to the “combi-triazene” SMA41, Cancer Chemother Pharmacol 51 (2003), pp. 11–20. Full Text via CrossRef | View Record in Scopus | Cited By in Scopus (19)
[21] Q. Qiu, F. Dudouit, S.L. Matheson, F. Brahimi, R. Banerjee and J.P. McNamee et al., The combi-targeting concept: a novel 3,3-disubstituted nitrosourea with EGFR tyrosine kinase inhibitory properties, Cancer Chemother Pharmacol 51 (2003), pp. 1–10. Full Text via CrossRef | View Record in Scopus | Cited By in Scopus (19)
[22] S. Matheson, J. McNamee and B.J. Jean-Claude, Design of a chimeric 3-methyl-1,2,3-triazene with mixed receptor tyrosine kinase and DNA damaging properties: a novel tumour targeting strategy, J Pharm Exp Ther 296 (2001), pp. 832–840. View Record in Scopus | Cited By in Scopus (29)
[23] K.D. Tew, L.C. Erickson, G. White, A.L. Wang, P.S. Schein and B. Hartley, Cytotoxicity of estramucine, a steroid-nitrogen mustard derivative, through non-DNA targets, Mol Pharmacol 24 (1983), pp. 324–328. View Record in Scopus | Cited By in Scopus (24)
[24] D. Vistica, P. Skehan, D. Scudiero, A. Monks, J.D. Pittam and M.R. Boyd, Tetrazolium-based assays for cellular viability: a critical examination of parameters affecting formazan production, Cancer Res 15 (1991), pp. 2515–2520. View Record in Scopus | Cited By in Scopus (193)
[25] J. Zimmermann, E. Buchdunger, H. Mett, T. Meyer, N. Lydon and B. Nicholas, Potent and selective inhibitors of the ABL-kinase: phenylaminopyrimidine (PAP) derivatives, Bioorg Med Chem Lett 7 (1997), pp. 187–192. Abstract | Abstract + References | PDF (279 K) | View Record in Scopus | Cited By in Scopus (87)
[26] M. Carroll, S. Ohno-Jones, S. Tamura, E. Buchdunger, J. Zimmermann and N.B. Lydon et al., CGP 57148, a tyrosine kinase inhibitor, inhibits the growth of cells expressing bcr-abl, tel-abl and tel-pdgfr, Blood 90 (1997), pp. 4947–4952.
[27] P. La Rosee, K. Johnson, A.S. Corbin, E.P. Stoffregen, E.M. Moseson and S. Willis et al., In vitro efficacy of combined treatment depends on the underlying mechanism of resistance in imatinib-resistant Bcr-Abl-positive cell lines, Blood 103 (2004), pp. 208–215. Full Text via CrossRef
[28] M. Lee, A.L. Rhodes, M.D. Wyatt, S. Forrow and J.A. Hartley, GC-base-sequence recognition by oligo(imidazolecarboxamide) and C-terminus-modified analogues of distamycin deduced from circular dichroism, proton nuclear magnetic resonance, and methidiumpropylethylenediaminetetraacetate-iron (II) footprinting studies, Biochemistry 32 (1993), pp. 4237–4245. Full Text via CrossRef | View Record in Scopus | Cited By in Scopus (58)
[29] M. Broggini, E. Erba, M. Ponti, D. Ballinari, C. Geroni and F. Spreafico et al., Selective DNA interaction of the novel distamycin derivative FCE 24517, Cancer Res 51 (1991), pp. 199–204. View Record in Scopus | Cited By in Scopus (64)
[30]
R. Aloyz, K. Grzywacz, Z.-Y. Xu, M. Loignon, M.A. Alaoui-Jamali and L. Panasci,
Imatinib sensitizes CLL lymphocytes to chlorambucil, Leukemia
18 (2004), pp. 409–414. Full Text via CrossRef | View Record in Scopus | Cited By in Scopus (9)
| Leukemia
Research Volume 29, Issue 5, May 2005, Pages 565-572 |
|
|
 |